Chapter 15 Regularization and Feature Selection
This section discusses a very common ML issue: you have too many features, and you do not know which ones to include. There are many ways to do feature selection, below we discuss a few more popular ones.
We assume you have loaded the following packages:
Below we load more as we introduce more.
For replicability, we also set the seed:
15.2 Forward selection
There are several solutions to this problem. A popular algorithm is forward selection where one first picks the best 1-feature model, thereafter tries adding all remaining features one-by-one to build the best two-feature model, and thereafter the best three-feature model, and so on, until the model performance starts to deteriorate.
15.2.1 Implement the algorithm manually
0-feature base model:
m0 = LinearRegression(fit_intercept = False)
X0 = np.ones((X.shape[0], 1))
X0t, X0v, yt, yv = train_test_split(X0, y, test_size = 0.4,
random_state = 8)
_t = m0.fit(X0t, yt)
m0.score(X0v, yv)## -0.006849647990198937Try 1-feature models:
best1 = {"i": -1, "R2": -np.inf}
m1 = LinearRegression(fit_intercept = True)
for i in range(X.shape[1]):
X1 = X.iloc[:,[i]]
X1t, X1v, yt, yv = train_test_split(X1, y, test_size = 0.4,
random_state = 8)
_t = m1.fit(X1t, yt)
s = m1.score(X1v, yv)
if s > best1["R2"]:
best1["i"] = i
best1["R2"] = s
best1## {'i': 9, 'R2': 0.4666428183761875}Add another feature:
best2 = {"i": -1, "R2": -np.inf}
m2 = LinearRegression(fit_intercept = True)
included_features = set([best1["i"]])
features_to_test = set(range(X.shape[1])) - included_features
for f in features_to_test:
features = included_features.union({f})
X2 = X.iloc[:, list(features)]
X2t, X2v, yt, yv = train_test_split(X2, y, test_size = 0.4,
random_state = 8)
_t = m2.fit(X2t, yt)
s = m2.score(X2v, yv)
if s > best2["R2"]:
best2["i"] = f
best2["R2"] = s
best2## {'i': 2, 'R2': 0.8677935861014889}Can add more in this fashion.
Make a function:
def bestFeature(X, y, included_features):
best = {"i": -1, "R2": -np.inf}
features_to_test = list(
set(range(X.shape[1])) - set(included_features))
for f in features_to_test:
features = included_features + [f]
Xf = X.iloc[:, features]
Xft, Xfv, yt, yv = train_test_split(Xf, y, test_size = 0.4,
random_state = 8)
_t = m2.fit(Xft, yt)
s = m2.score(Xfv, yv)
if s > best["R2"]:
best["i"] = f
best["R2"] = s
return best
best = list()
included_features = []
r2s = []
for f in range(X.shape[1]):
b = bestFeature(X, y, included_features)
best.append(b)
included_features = included_features + [b["i"]]
r2s.append(b["R2"])
best## [{'i': 9, 'R2': 0.4666428183761875}, {'i': 2, 'R2': 0.8677935861014889}, {'i': 27, 'R2': 0.9296522813032738}, {'i': 28, 'R2': 0.9396723227644169}, {'i': 14, 'R2': 0.9430611422142994}, {'i': 22, 'R2': 0.9437993501430608}, {'i': 33, 'R2': 0.9446022123630592}, {'i': 59, 'R2': 0.9450369454683462}, {'i': 10, 'R2': 0.9452498568205782}, {'i': 8, 'R2': 0.9455986815032276}, {'i': 19, 'R2': 0.9462862467428159}, {'i': 1, 'R2': 0.9464320773146165}, {'i': 46, 'R2': 0.9465914853888853}, {'i': 40, 'R2': 0.9465889082229213}, {'i': 55, 'R2': 0.9465359196145449}, {'i': 36, 'R2': 0.9464542982901596}, {'i': 5, 'R2': 0.9463762822853345}, {'i': 20, 'R2': 0.9462833979835922}, {'i': 34, 'R2': 0.9462050456158057}, {'i': 48, 'R2': 0.9461478229257206}, {'i': 50, 'R2': 0.9461054197700751}, {'i': 52, 'R2': 0.9460730478692239}, {'i': 26, 'R2': 0.9459904691472554}, {'i': 42, 'R2': 0.9459265694540916}, {'i': 44, 'R2': 0.9458087497124463}, {'i': 32, 'R2': 0.9455590142220006}, {'i': 39, 'R2': 0.945408502141411}, {'i': 25, 'R2': 0.9449675269753557}, {'i': 24, 'R2': 0.9445346914386826}, {'i': 58, 'R2': 0.9438665545550707}, {'i': 54, 'R2': 0.9431161004717253}, {'i': 38, 'R2': 0.9425908604390918}, {'i': 21, 'R2': 0.9418808150038418}, {'i': 35, 'R2': 0.9414073375383587}, {'i': 37, 'R2': 0.9410776069690762}, {'i': 41, 'R2': 0.940836636399586}, {'i': 49, 'R2': 0.9406534112839524}, {'i': 51, 'R2': 0.940509615245991}, {'i': 53, 'R2': 0.9403938540834724}, {'i': 43, 'R2': 0.9401119255355864}, {'i': 57, 'R2': 0.9397885757150655}, {'i': 23, 'R2': 0.9394085387576068}, {'i': 12, 'R2': 0.9379313435724337}, {'i': 30, 'R2': 0.9377835490332745}, {'i': 45, 'R2': 0.9333091167915868}, {'i': 29, 'R2': 0.9340676090843362}, {'i': 16, 'R2': 0.9285874234675286}, {'i': 0, 'R2': 0.9182942310441039}, {'i': 6, 'R2': 0.880809369532485}, {'i': 56, 'R2': 0.8437760019671372}, {'i': 31, 'R2': 0.8474899063079687}, {'i': 47, 'R2': 0.8521445384395225}, {'i': 17, 'R2': -9.818297650553768e+22}, {'i': 13, 'R2': -2.000001247173613e+22}, {'i': 15, 'R2': -4.135131612681403e+22}, {'i': 3, 'R2': -2.7600188239712023e+22}, {'i': 4, 'R2': -1.1141317933826631e+23}, {'i': 7, 'R2': -5.35812572220869e+22}, {'i': 18, 'R2': -3.2569968014607136e+22}, {'i': 11, 'R2': -3.4476870254964047e+22}]## (-0.1, 1.0)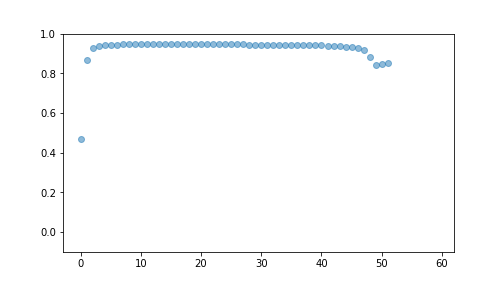
plot of chunk unnamed-chunk-12
15.2.2 sklearn’s SequentialFeatureSelector
When using sklearn, you can do forward and backward selection using
SequentialFeatureSelector(). The usage of it is fairly similar to
that of any sklearn model. First you need to make the
SequentialFeatureSelector-object. It’s most important argument is the
model, direction (“forward” or “backward”) scoring (whether to use
\(R^2\) or another model goodness measure), and the number of
cross-validation folds. Here is a small example:
from sklearn.feature_selection import SequentialFeatureSelector
m = LinearRegression()
sfs = SequentialFeatureSelector(m, direction = "forward")
_t = sfs.fit(Xt, yt)The .fit() method runs the algorithm. This may be rather slow,
depending on your data size.
After fitting, you can query the included features:
## array(['x3', 'x10', 'z1_B', 'z1_C', 'z2_B', 'z2_C', 'z4_B', 'z4_C',
## 'z5_B', 'z6_B', 'z7_B', 'z7_C', 'z8_B', 'z10_B', 'z10_C', 'z12_B',
## 'z12_C', 'z13_B', 'z13_C', 'z14_B', 'z14_C', 'z15_B', 'z15_C',
## 'z16_B', 'z17_B', 'z18_B', 'z18_C', 'z19_B', 'z20_B', 'z20_C'],
## dtype=object)As you can see, forward selection included only two x-s but many z-s.
You can extract only the included features by selecting the names:
## (60, 30)or by using the .transform() method:
As you see, forward selection only preserved 30 columns.
Finally, the validation \(R^2\) is
## 0.9329183739713025The validation \(R^2\) is much larger than when using all the features in Section 15.1.
Exercise 15.2 Replicate the above with backward selection. Do you get similar features? Do you get similar \(R^2\)?
15.3 Ridge and Lasso regression
Ridge and Lasso are methods that are related to forward selection. These methods penalize large \(\beta\) values and hence suppress or eliminate correlated variables. These do not need looping over different combinations of variables like forward selection, however, one normally has to loop over the penalty parameter alpha to find the optimal value.
Both methods are living in sklearn.linear_regression.
We demonstrate this with ridge regression below:
from sklearn.linear_model import Ridge, Lasso
from sklearn.model_selection import cross_val_score
m = Ridge(alpha = 1) # penalty value 1
cross_val_score(m, X, y, cv=10).mean()## 0.908232518044392## 0.9035580051705517As evident from the above, penalty values 1 and 0.3 give virtually equal results. The cross-validated \(R^2\) is also comparable to what forward-selection suggested.
Lasso works in an analogous fashion as ridge, but as it’s penalty is not globally differentiable (it is \(L_1\) penalty), then you may run into more problems with convergence.
Both methods have more options, in particular they can normalize the features before fitting.
## 0.9264571331356263## 0.8627943726880424