EZLearn+Automatic Claim Validation
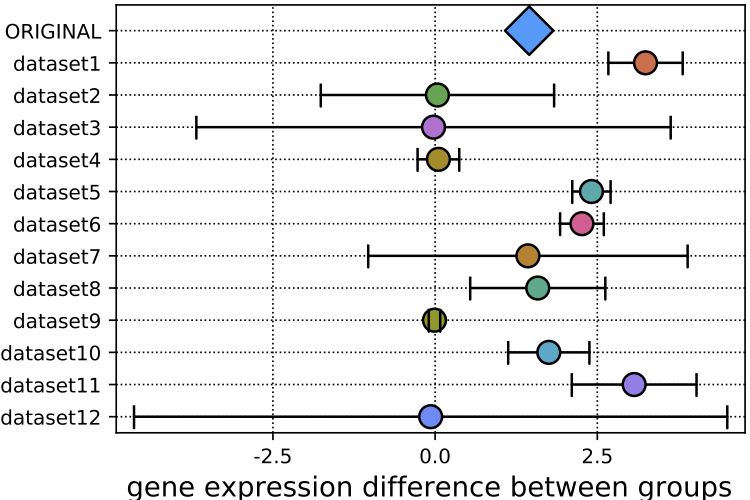
We combine distant supervision and co-learning to classify scientific datasets with no training data. The method, which we refer to as EZLearn can handle potentially thousands of classes that may change frequently, such that supervised approaches become infeasible.
The key insight is to leverage the abundant unlabeled data together with two sources of ``organic’’ supervision: a lexicon for the annotation classes and free-text descriptions that often accompany unlabeled data. Such indirect supervision is readily available in science and other high-value applications.
To exploit these sources of information, EZLearn introduces an auxiliary natural language processing system, which uses the lexicon to generate initial noisy labels from the text descriptions, and then co-teaches the main classifier until convergence. Effectively, EZLearncombines distant supervision and co-training into a new learning paradigm for leveraging unlabeled data. Because no hand-labeled examples are required, EZLearn is naturally applicable to domains with a long tail of classes and/or frequent updates.
We evaluated EZLearn on applications in functional genomics and scientific figure comprehension. In both cases, using text descriptions as the pivot, EZLearn learned to accurately annotate data samples without direct supervision, even substantially outperforming the state-of-the-art supervised methods trained on tens of thousands of annotated examples.
EZLearn is one component of a broader agenda to automatically validate scientific claims against published data. Researchers are increasingly required to submit data to public repositories as a condition of publication or funding, but this data remains underused and concerns about reproducibility continue to mount. We are developing an end-to-end system that can capture scientific claims from papers, validate them against the data associated with the paper, then generalize and adapt the claims to other relevant datasets in the repository to gather additional statistical evidence.
Collaborators
- Hoifung Poon, Microsoft Research
Students
- Maxim Grechkin
Publications
- EZLearn: Exploiting Organic Supervision in Large-Scale Data Annotation
Maxim Grechkin, Hoifung Poon, Bill Howe.
Learning with Limited Labeled Data Workshop (co-located with NIPS 2017) (best paper award) 2017@article{grechkin17ezlearn, author = {Grechkin, Maxim and Poon, Hoifung and Howe, Bill}, title = {EZLearn: Exploiting Organic Supervision in Large-Scale Data Annotation}, journal = {Learning with Limited Labeled Data Workshop (co-located with NIPS 2017) (best paper award)}, year = {2017}, url = {http://arxiv.org/abs/1709.08600} }
- Wide-Open: Accelerating public data release by automating detection of overdue datasets
Maxim Grechkin, Hoifung Poon, Bill Howe.
PLoS biology 15(6) 2017@article{grechkin2017wide, title = {Wide-Open: Accelerating public data release by automating detection of overdue datasets}, author = {Grechkin, Maxim and Poon, Hoifung and Howe, Bill}, journal = {PLoS biology}, volume = {15}, number = {6}, pages = {e2002477}, year = {2017}, publisher = {Public Library of Science} }
Sponsors
projects
This webpage was built with Bootstrap and Jekyll. You can find the source code here. Last updated: Aug 02, 2021