Scalable Flow Cytometry
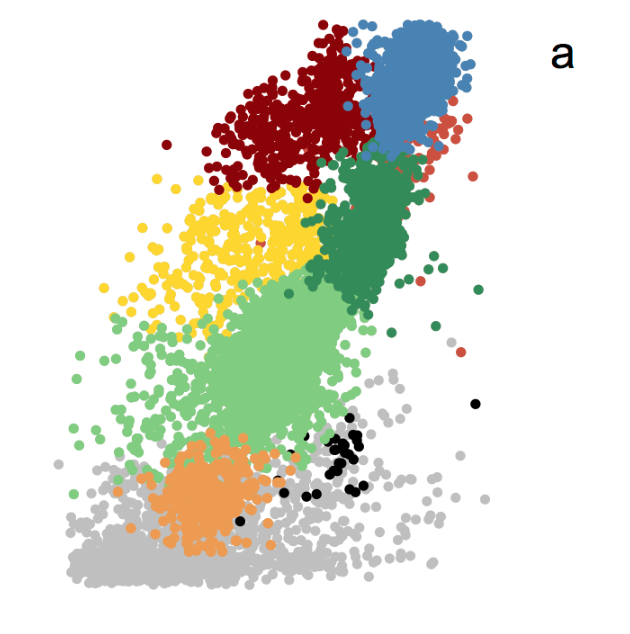
Recent technological innovations in flow cytometry now allow oceanographers to collect high-frequency flow cytometry data from particles in aquatic environments on a scale far surpassing conventional flow cytometers. The SeaFlow cytometer continuously profiles microbial phytoplankton populations across thousands of kilometers of the surface ocean. The data streams produced by instruments such as SeaFlow challenge the traditional sample-by- sample approach in cytometric analysis and illuminate the need for scalable clustering algorithms to extract population information from these large-scale, high-frequency flow cytometers.
We explore how available algorithms commonly used for medical applications perform at classification of such a large-scale, environmental flow cytometry data. We apply large-scale Gaussian mixture models (GMM) to massive data sets using Hadoop. This approach outperforms current state-of-the-art cytometry classification algorithms in accuracy, and can be coupled with manual or automatic partitioning of data into homogeneous sections for further classification gains. We propose the GMM with partitioning approach for classification of large-scale, high-frequency flow cytometry data.
In earlier work, we developed systems for managing ad hoc sensor data, recommending visualizations, and managing complex workflows all motivated in part by SeaFlow.
People
- Dan Halperin, lead
- Jeremy Hyrkas
- Shrainik Jain
- Sagar Chitnis
Papers
- Scalable clustering algorithms for continuous environmental flow cytometry
Jeremy Hyrkas, Sophie Clayton, Francois Ribalet, Daniel Halperin, E. Virginia Armbrust, Bill Howe.
Bioinformatics 32(3) 2016@article{hyrkas2016clustering, author = {Hyrkas, Jeremy and Clayton, Sophie and Ribalet, Francois and Halperin, Daniel and Armbrust, E. Virginia and Howe, Bill}, title = {Scalable clustering algorithms for continuous environmental flow cytometry}, journal = {Bioinformatics}, volume = {32}, number = {3}, pages = {417--423}, year = {2016}, url = {http://dx.doi.org/10.1093/bioinformatics/btv594}, doi = {10.1093/bioinformatics/btv594}, timestamp = {Fri, 05 Feb 2016 14:35:15 +0100}, biburl = {http://dblp.uni-trier.de/rec/bib/journals/bioinformatics/HyrkasCRHAH16}, bibsource = {dblp computer science bibliography, http://dblp.org} }
- Time-varying clusters in large-scale flow cytometry
Jeremy Hyrkas, Daniel Halperin, Bill Howe.
IAAI Conference 2015@inproceedings{hyrkas2015time, title = {Time-varying clusters in large-scale flow cytometry}, author = {Hyrkas, Jeremy and Halperin, Daniel and Howe, Bill}, booktitle = {IAAI Conference}, year = {2015} }
- Gaussian mixture models use-case: in-memory analysis with myria
Ryan Maas, Jeremy Hyrkas, Olivia Grace Telford, Magdalena Balazinska, Andrew Connolly, Bill Howe.
Proceedings of the 3rd VLDB Workshop on In-Memory Data Mangement and Analytics 2015@inproceedings{maas2015gaussian, title = {Gaussian mixture models use-case: in-memory analysis with myria}, author = {Maas, Ryan and Hyrkas, Jeremy and Telford, Olivia Grace and Balazinska, Magdalena and Connolly, Andrew and Howe, Bill}, booktitle = {Proceedings of the 3rd VLDB Workshop on In-Memory Data Mangement and Analytics}, pages = {3}, year = {2015}, organization = {ACM} }
- Collaborative science workflows in SQL
Bill Howe, Daniel Halperin, Francois Ribalet, Sagar Chitnis, E Virginia Armbrust.
Computing in Science and Engineering 15(3) 2013@article{howe2013collaborative, title = {Collaborative science workflows in SQL}, author = {Howe, Bill and Halperin, Daniel and Ribalet, Francois and Chitnis, Sagar and Armbrust, E Virginia}, journal = {Computing in Science and Engineering}, volume = {15}, number = {3}, pages = {22--31}, year = {2013}, publisher = {AIP Publishing} }
- SQLShare: Scientific workflow via relational view sharing
Bill Howe, Francois Ribalet, Daniel Halperin, Sagar Chitnis, E Virginia Armbrust.
Computing in Science and Engineering, Special Issue on Science Data Management 15(2) 2013
projects
This webpage was built with Bootstrap and Jekyll. You can find the source code here. Last updated: Aug 02, 2021