The Morris Lab
RNA is a mutlifaceted molecule - informational, catalytic, regulatory,
structural - but its dominant role from the standpoint of cell
phenotype is to be translated into protein. Regulation of gene
expression at the level of translation provides a massive and rapid
way for a cell to respond to physical, chemical and biological
changes in its environment. We study the regulation and
coordination of mRNA translation both at the genome-wide scale
and at the level of individual molecules.
NEWS
Interrogating the Translated
Transcriptome
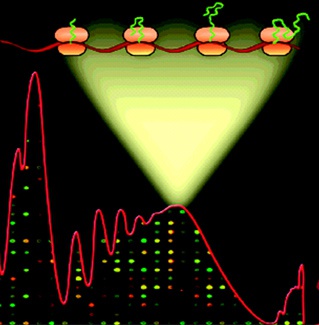
TSAA. Traditional approaches to genome-wide expression analysis measure total transcript levels.
This is valuable information, but it gives no insight into the rates of synthesis of the proteins encoded
by a transcriptome. We developed Translation State Array Analysis or TSAA, which allows an
estimate of the loading of ribosomes onto the mRNAs of a transcriptome as well as transcript levels.
We have used this approach to interrogate several biological systems, including responses of yeast to
mating pheromone and cellular stresses, gene expression in differentiating ES cells and translational
control upon macrophage attachment to a substratum. See Morris (2009) for a discussion of an
exciting new approach to exploration of translation state.
RiboTag. A major issue in genome biology is how to examine gene expression in the individual cells
in a tissue. The RiboTag mouse provides a powerful tool for interrogating the ribosome-associated
transcriptomes of single cell types in complex tissues such as brain. The approach is readily applied to
any cell type for which a specific Cre recombinase driver is available.